Title
Usage
runChASM(
rawReadCountsIn,
minSamplesPerProtocol = 30,
min_reads = 60000,
max_reads = 1e+09,
p_contamination = 0.01,
show_plot = TRUE,
printMissingIDs = FALSE
)Arguments
- rawReadCountsIn
the reads counts for each chromosome
- minSamplesPerProtocol
minimum number of reads per protocol for parameter estimation
- min_reads
the minimum number of reads for Dirichlet parameter estimation
- max_reads
the maximum number of reads for Dirichlet parameter estimation
- p_contamination
the probability of a sample yielding significant contamination
- show_plot
show the clustering plot for sex chromosomal aneuploidy Dirichlet estimation?
- printMissingIDs
when combining karyotype calls, return names that are missing (or just the number of missing IDs)?
Examples
runChASM(rawReadCountsIn = example_data)
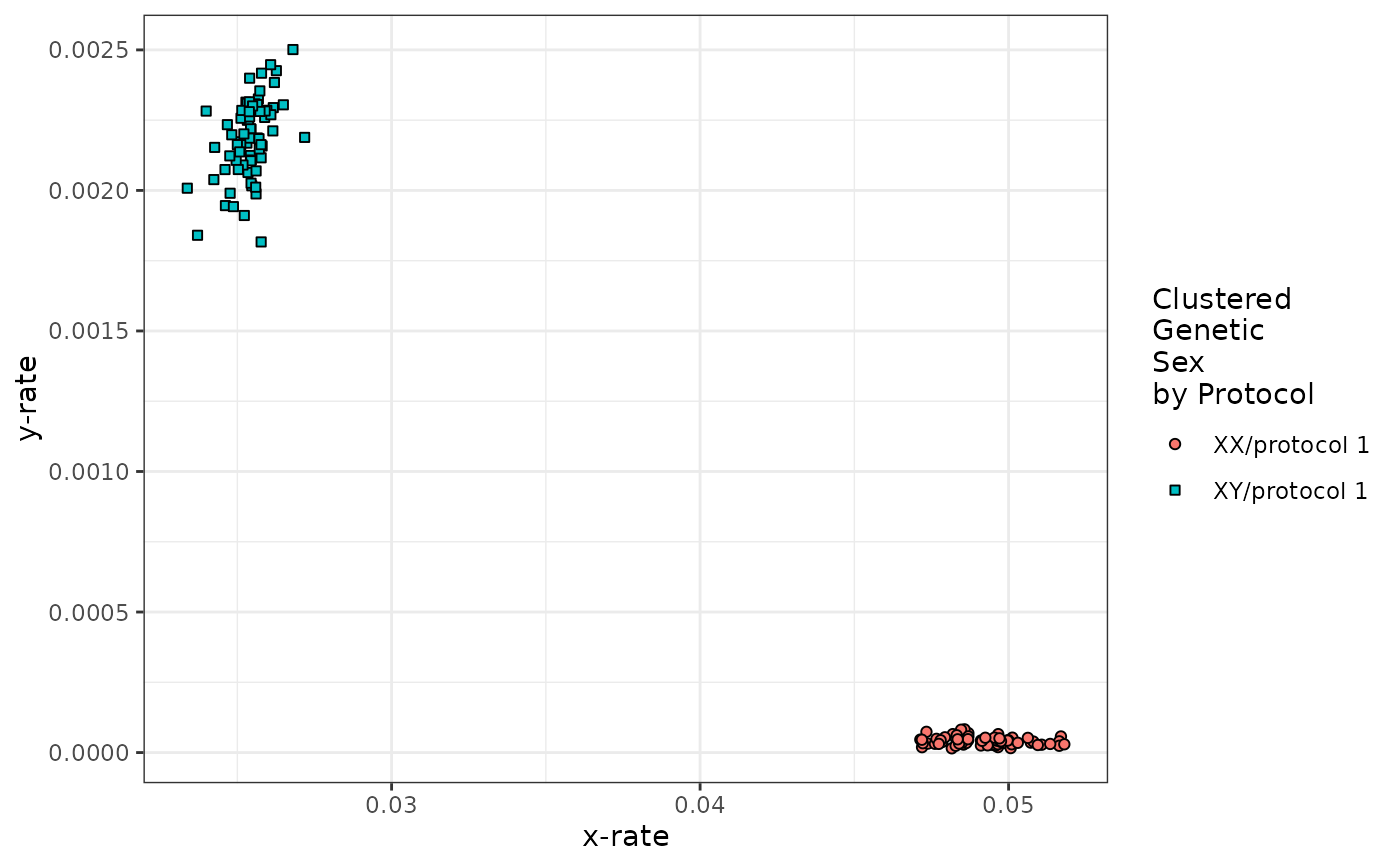 #> $karyotypes
#> # A tibble: 222 × 11
#> sample protocol unusual flags autosomal_call sca_call C_call autosomal_total
#> <chr> <chr> <lgl> <int> <chr> <chr> <chr> <dbl>
#> 1 Ind_1_1 protoco… TRUE 3 No Aneuploidy XX No Si… 89539
#> 2 Ind_1_2 protoco… FALSE 1 No Aneuploidy XX No Si… 48376
#> 3 Ind_3_1 protoco… TRUE 2 No Aneuploidy XY No Si… 113335
#> 4 Ind_4_1 protoco… FALSE 0 No Aneuploidy XX No Si… 406472
#> 5 Ind_5_1 protoco… FALSE 0 No Aneuploidy XX No Si… 27731
#> 6 Ind_6_1 protoco… FALSE 0 No Aneuploidy XY No Si… 681256
#> 7 Ind_7_2 protoco… FALSE 0 No Aneuploidy XY No Si… 130928
#> 8 Ind_9_1 protoco… FALSE 0 No Aneuploidy XX No Si… 425906
#> 9 Ind_15… protoco… FALSE 1 No Aneuploidy XX No Si… 3364
#> 10 Ind_17… protoco… FALSE 1 No Aneuploidy XY No Si… 11699
#> # ℹ 212 more rows
#> # ℹ 3 more variables: sca_total <dbl>, automsomal_maxP <dbl>, sca_maxP <dbl>
#>
#> $karyotypes.auto
#> # A tibble: 222 × 81
#> sample total P_call maxP protocol chr1 chr2 chr3 chr4 chr5 chr6 chr7
#> <chr> <dbl> <chr> <dbl> <chr> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 Ind_1… 89539 No An… 1 protoco… 7471 7955 6581 6144 5983 5704 5084
#> 2 Ind_1… 48376 No An… 1 protoco… 4248 4206 3367 2968 3039 2888 2735
#> 3 Ind_3… 113335 No An… 1 protoco… 9839 9935 7947 6918 7094 6847 6300
#> 4 Ind_4… 406472 No An… 1 protoco… 34248 35719 29423 26958 26557 25109 22975
#> 5 Ind_5… 27731 No An… 1.000 protoco… 2306 2357 2029 1791 1813 1657 1576
#> 6 Ind_6… 681256 No An… 1 protoco… 58317 59471 48482 43163 43211 41308 38218
#> 7 Ind_7… 130928 No An… 1 protoco… 11098 11528 9380 8495 8376 8117 7300
#> 8 Ind_9… 425906 No An… 1 protoco… 36039 37919 30481 27936 27719 26142 23895
#> 9 Ind_1… 3364 No An… 1.000 protoco… 298 297 241 239 227 211 198
#> 10 Ind_1… 11699 No An… 1.000 protoco… 999 1054 798 745 776 726 636
#> # ℹ 212 more rows
#> # ℹ 69 more variables: chr8 <dbl>, chr9 <dbl>, chr10 <dbl>, chr11 <dbl>,
#> # chr12 <dbl>, chr13 <dbl>, chr14 <dbl>, chr15 <dbl>, chr16 <dbl>,
#> # chr17 <dbl>, chr18 <dbl>, chr19 <dbl>, chr20 <dbl>, chr21 <dbl>,
#> # chr22 <dbl>, p1 <dbl>, p2 <dbl>, p3 <dbl>, p4 <dbl>, p5 <dbl>, p6 <dbl>,
#> # p7 <dbl>, p8 <dbl>, p9 <dbl>, p10 <dbl>, p11 <dbl>, p12 <dbl>, p13 <dbl>,
#> # p14 <dbl>, p15 <dbl>, p16 <dbl>, p17 <dbl>, p18 <dbl>, p19 <dbl>, …
#>
#> $karyotypes.sca
#> # A tibble: 222 × 34
#> # Rowwise:
#> sample protocol total P_call maxP P_cont auto X Y px py
#> <chr> <chr> <dbl> <chr> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 Ind_1_1 protoco… 94269 XX 1 -39.2 89539 4725 5 0.0501 5.30e-5
#> 2 Ind_1_2 protoco… 50760 XX 1 -26.0 48376 2382 2 0.0469 3.94e-5
#> 3 Ind_3_1 protoco… 116538 XY 1 -29.4 113335 2968 235 0.0255 2.02e-3
#> 4 Ind_4_1 protoco… 427738 XX 1 -41.6 406472 21254 12 0.0497 2.81e-5
#> 5 Ind_5_1 protoco… 29216 XX 1 -36.4 27731 1484 1 0.0508 3.42e-5
#> 6 Ind_6_1 protoco… 700574 XY 1 -33.0 681256 17742 1576 0.0253 2.25e-3
#> 7 Ind_7_2 protoco… 134684 XY 1 -29.7 130928 3471 285 0.0258 2.12e-3
#> 8 Ind_9_1 protoco… 448296 XX 1 -41.8 425906 22375 15 0.0499 3.35e-5
#> 9 Ind_15… protoco… 3530 XX 1.000 -14.3 3364 166 0 0.0470 0
#> 10 Ind_17… protoco… 12023 XY 1.000 -24.9 11699 298 26 0.0248 2.16e-3
#> # ℹ 212 more rows
#> # ℹ 23 more variables: pz <dbl>, ax <dbl>, ay <dbl>, az <dbl>, a0 <dbl>,
#> # correction <dbl>, Gamma <dbl>, P_XY <dbl>, P_XX <dbl>, P_XXY <dbl>,
#> # P_X <dbl>, P_XXX <dbl>, P_XYY <dbl>, SumP <dbl>, SumC <dbl>,
#> # Posterior_XY <dbl>, Posterior_XX <dbl>, Posterior_XXY <dbl>,
#> # Posterior_X <dbl>, Posterior_XXX <dbl>, Posterior_XYY <dbl>,
#> # Posterior_cont <dbl>, P_C <chr>
#>
#> $dirichlet.auto
#> # A tibble: 1 × 24
#> protocol a1 a2 a3 a4 a5 a6 a7 a8 a9 a10 a11
#> <chr> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 protocol 1 3770. 3914. 3195. 2901. 2853. 2732. 2489. 2362. 1870. 2226. 2224.
#> # ℹ 12 more variables: a12 <dbl>, a13 <dbl>, a14 <dbl>, a15 <dbl>, a16 <dbl>,
#> # a17 <dbl>, a18 <dbl>, a19 <dbl>, a20 <dbl>, a21 <dbl>, a22 <dbl>, a0 <dbl>
#>
#> $dirichlet.sca
#> # A tibble: 1 × 6
#> protocol ax ay az a0 correction
#> <fct> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 protocol 1 1984. 172. 75939. 78094. 0.0000581
#>
#> $z.scores
#> # A tibble: 4,884 × 15
#> sample protocol chr Nij Nj alpha alpha0 muij sigmaij Zij flag
#> <chr> <chr> <chr> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <lgl>
#> 1 Ind_10… protoco… chr1 60482 710804 3770. 44487. 60240. 967. 0.250 FALSE
#> 2 Ind_10… protoco… chr2 62847 710804 3914. 44487. 62530. 984. 0.322 FALSE
#> 3 Ind_10… protoco… chr3 51218 710804 3195. 44487. 51052. 897. 0.185 FALSE
#> 4 Ind_10… protoco… chr4 46642 710804 2901. 44487. 46354. 858. 0.336 FALSE
#> 5 Ind_10… protoco… chr5 45710 710804 2853. 44487. 45577. 851. 0.156 FALSE
#> 6 Ind_10… protoco… chr6 43676 710804 2732. 44487. 43653. 834. 0.0281 FALSE
#> 7 Ind_10… protoco… chr7 39802 710804 2489. 44487. 39776. 798. 0.0330 FALSE
#> 8 Ind_10… protoco… chr8 37495 710804 2362. 44487. 37740. 779. -0.315 FALSE
#> 9 Ind_10… protoco… chr9 30034 710804 1870. 44487. 29871. 697. 0.234 FALSE
#> 10 Ind_10… protoco… chr10 35418 710804 2226. 44487. 35561. 757. -0.189 FALSE
#> # ℹ 4,874 more rows
#> # ℹ 4 more variables: Xj <dbl>, p <dbl>, flags <int>, unusual <lgl>
#>
#> $minSamplesPerProtocol
#> [1] 30
#>
#> $min_reads
#> [1] 60000
#>
#> $max_reads
#> [1] 1e+09
#>
#> $p_contamination
#> [1] 0.01
#>
#> $karyotypes
#> # A tibble: 222 × 11
#> sample protocol unusual flags autosomal_call sca_call C_call autosomal_total
#> <chr> <chr> <lgl> <int> <chr> <chr> <chr> <dbl>
#> 1 Ind_1_1 protoco… TRUE 3 No Aneuploidy XX No Si… 89539
#> 2 Ind_1_2 protoco… FALSE 1 No Aneuploidy XX No Si… 48376
#> 3 Ind_3_1 protoco… TRUE 2 No Aneuploidy XY No Si… 113335
#> 4 Ind_4_1 protoco… FALSE 0 No Aneuploidy XX No Si… 406472
#> 5 Ind_5_1 protoco… FALSE 0 No Aneuploidy XX No Si… 27731
#> 6 Ind_6_1 protoco… FALSE 0 No Aneuploidy XY No Si… 681256
#> 7 Ind_7_2 protoco… FALSE 0 No Aneuploidy XY No Si… 130928
#> 8 Ind_9_1 protoco… FALSE 0 No Aneuploidy XX No Si… 425906
#> 9 Ind_15… protoco… FALSE 1 No Aneuploidy XX No Si… 3364
#> 10 Ind_17… protoco… FALSE 1 No Aneuploidy XY No Si… 11699
#> # ℹ 212 more rows
#> # ℹ 3 more variables: sca_total <dbl>, automsomal_maxP <dbl>, sca_maxP <dbl>
#>
#> $karyotypes.auto
#> # A tibble: 222 × 81
#> sample total P_call maxP protocol chr1 chr2 chr3 chr4 chr5 chr6 chr7
#> <chr> <dbl> <chr> <dbl> <chr> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 Ind_1… 89539 No An… 1 protoco… 7471 7955 6581 6144 5983 5704 5084
#> 2 Ind_1… 48376 No An… 1 protoco… 4248 4206 3367 2968 3039 2888 2735
#> 3 Ind_3… 113335 No An… 1 protoco… 9839 9935 7947 6918 7094 6847 6300
#> 4 Ind_4… 406472 No An… 1 protoco… 34248 35719 29423 26958 26557 25109 22975
#> 5 Ind_5… 27731 No An… 1.000 protoco… 2306 2357 2029 1791 1813 1657 1576
#> 6 Ind_6… 681256 No An… 1 protoco… 58317 59471 48482 43163 43211 41308 38218
#> 7 Ind_7… 130928 No An… 1 protoco… 11098 11528 9380 8495 8376 8117 7300
#> 8 Ind_9… 425906 No An… 1 protoco… 36039 37919 30481 27936 27719 26142 23895
#> 9 Ind_1… 3364 No An… 1.000 protoco… 298 297 241 239 227 211 198
#> 10 Ind_1… 11699 No An… 1.000 protoco… 999 1054 798 745 776 726 636
#> # ℹ 212 more rows
#> # ℹ 69 more variables: chr8 <dbl>, chr9 <dbl>, chr10 <dbl>, chr11 <dbl>,
#> # chr12 <dbl>, chr13 <dbl>, chr14 <dbl>, chr15 <dbl>, chr16 <dbl>,
#> # chr17 <dbl>, chr18 <dbl>, chr19 <dbl>, chr20 <dbl>, chr21 <dbl>,
#> # chr22 <dbl>, p1 <dbl>, p2 <dbl>, p3 <dbl>, p4 <dbl>, p5 <dbl>, p6 <dbl>,
#> # p7 <dbl>, p8 <dbl>, p9 <dbl>, p10 <dbl>, p11 <dbl>, p12 <dbl>, p13 <dbl>,
#> # p14 <dbl>, p15 <dbl>, p16 <dbl>, p17 <dbl>, p18 <dbl>, p19 <dbl>, …
#>
#> $karyotypes.sca
#> # A tibble: 222 × 34
#> # Rowwise:
#> sample protocol total P_call maxP P_cont auto X Y px py
#> <chr> <chr> <dbl> <chr> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 Ind_1_1 protoco… 94269 XX 1 -39.2 89539 4725 5 0.0501 5.30e-5
#> 2 Ind_1_2 protoco… 50760 XX 1 -26.0 48376 2382 2 0.0469 3.94e-5
#> 3 Ind_3_1 protoco… 116538 XY 1 -29.4 113335 2968 235 0.0255 2.02e-3
#> 4 Ind_4_1 protoco… 427738 XX 1 -41.6 406472 21254 12 0.0497 2.81e-5
#> 5 Ind_5_1 protoco… 29216 XX 1 -36.4 27731 1484 1 0.0508 3.42e-5
#> 6 Ind_6_1 protoco… 700574 XY 1 -33.0 681256 17742 1576 0.0253 2.25e-3
#> 7 Ind_7_2 protoco… 134684 XY 1 -29.7 130928 3471 285 0.0258 2.12e-3
#> 8 Ind_9_1 protoco… 448296 XX 1 -41.8 425906 22375 15 0.0499 3.35e-5
#> 9 Ind_15… protoco… 3530 XX 1.000 -14.3 3364 166 0 0.0470 0
#> 10 Ind_17… protoco… 12023 XY 1.000 -24.9 11699 298 26 0.0248 2.16e-3
#> # ℹ 212 more rows
#> # ℹ 23 more variables: pz <dbl>, ax <dbl>, ay <dbl>, az <dbl>, a0 <dbl>,
#> # correction <dbl>, Gamma <dbl>, P_XY <dbl>, P_XX <dbl>, P_XXY <dbl>,
#> # P_X <dbl>, P_XXX <dbl>, P_XYY <dbl>, SumP <dbl>, SumC <dbl>,
#> # Posterior_XY <dbl>, Posterior_XX <dbl>, Posterior_XXY <dbl>,
#> # Posterior_X <dbl>, Posterior_XXX <dbl>, Posterior_XYY <dbl>,
#> # Posterior_cont <dbl>, P_C <chr>
#>
#> $dirichlet.auto
#> # A tibble: 1 × 24
#> protocol a1 a2 a3 a4 a5 a6 a7 a8 a9 a10 a11
#> <chr> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 protocol 1 3770. 3914. 3195. 2901. 2853. 2732. 2489. 2362. 1870. 2226. 2224.
#> # ℹ 12 more variables: a12 <dbl>, a13 <dbl>, a14 <dbl>, a15 <dbl>, a16 <dbl>,
#> # a17 <dbl>, a18 <dbl>, a19 <dbl>, a20 <dbl>, a21 <dbl>, a22 <dbl>, a0 <dbl>
#>
#> $dirichlet.sca
#> # A tibble: 1 × 6
#> protocol ax ay az a0 correction
#> <fct> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 protocol 1 1984. 172. 75939. 78094. 0.0000581
#>
#> $z.scores
#> # A tibble: 4,884 × 15
#> sample protocol chr Nij Nj alpha alpha0 muij sigmaij Zij flag
#> <chr> <chr> <chr> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <lgl>
#> 1 Ind_10… protoco… chr1 60482 710804 3770. 44487. 60240. 967. 0.250 FALSE
#> 2 Ind_10… protoco… chr2 62847 710804 3914. 44487. 62530. 984. 0.322 FALSE
#> 3 Ind_10… protoco… chr3 51218 710804 3195. 44487. 51052. 897. 0.185 FALSE
#> 4 Ind_10… protoco… chr4 46642 710804 2901. 44487. 46354. 858. 0.336 FALSE
#> 5 Ind_10… protoco… chr5 45710 710804 2853. 44487. 45577. 851. 0.156 FALSE
#> 6 Ind_10… protoco… chr6 43676 710804 2732. 44487. 43653. 834. 0.0281 FALSE
#> 7 Ind_10… protoco… chr7 39802 710804 2489. 44487. 39776. 798. 0.0330 FALSE
#> 8 Ind_10… protoco… chr8 37495 710804 2362. 44487. 37740. 779. -0.315 FALSE
#> 9 Ind_10… protoco… chr9 30034 710804 1870. 44487. 29871. 697. 0.234 FALSE
#> 10 Ind_10… protoco… chr10 35418 710804 2226. 44487. 35561. 757. -0.189 FALSE
#> # ℹ 4,874 more rows
#> # ℹ 4 more variables: Xj <dbl>, p <dbl>, flags <int>, unusual <lgl>
#>
#> $minSamplesPerProtocol
#> [1] 30
#>
#> $min_reads
#> [1] 60000
#>
#> $max_reads
#> [1] 1e+09
#>
#> $p_contamination
#> [1] 0.01
#>