Final version is based on BR's version
Usage
plot_ergm_bic(
ergms,
text_size = 10,
text_angle = 90,
abbr = FALSE,
measure = "BIC",
top_5 = FALSE
)Examples
ergms <- example_network |>
get_ergms(
preds = c("site", "genetic_sex"),
types = c("nodematch", "nodemix"))
#> Starting maximum pseudolikelihood estimation (MPLE):
#> Obtaining the responsible dyads.
#> Evaluating the predictor and response matrix.
#> Maximizing the pseudolikelihood.
#> Finished MPLE.
#> Evaluating log-likelihood at the estimate.
#>
#> Starting maximum pseudolikelihood estimation (MPLE):
#> Obtaining the responsible dyads.
#> Evaluating the predictor and response matrix.
#> Maximizing the pseudolikelihood.
#> Finished MPLE.
#> Evaluating log-likelihood at the estimate.
#>
#> Starting maximum pseudolikelihood estimation (MPLE):
#> Obtaining the responsible dyads.
#> Evaluating the predictor and response matrix.
#> Maximizing the pseudolikelihood.
#> Finished MPLE.
#> Evaluating log-likelihood at the estimate.
#>
#> Starting maximum pseudolikelihood estimation (MPLE):
#> Obtaining the responsible dyads.
#> Evaluating the predictor and response matrix.
#> Maximizing the pseudolikelihood.
#> Finished MPLE.
#> Evaluating log-likelihood at the estimate.
#>
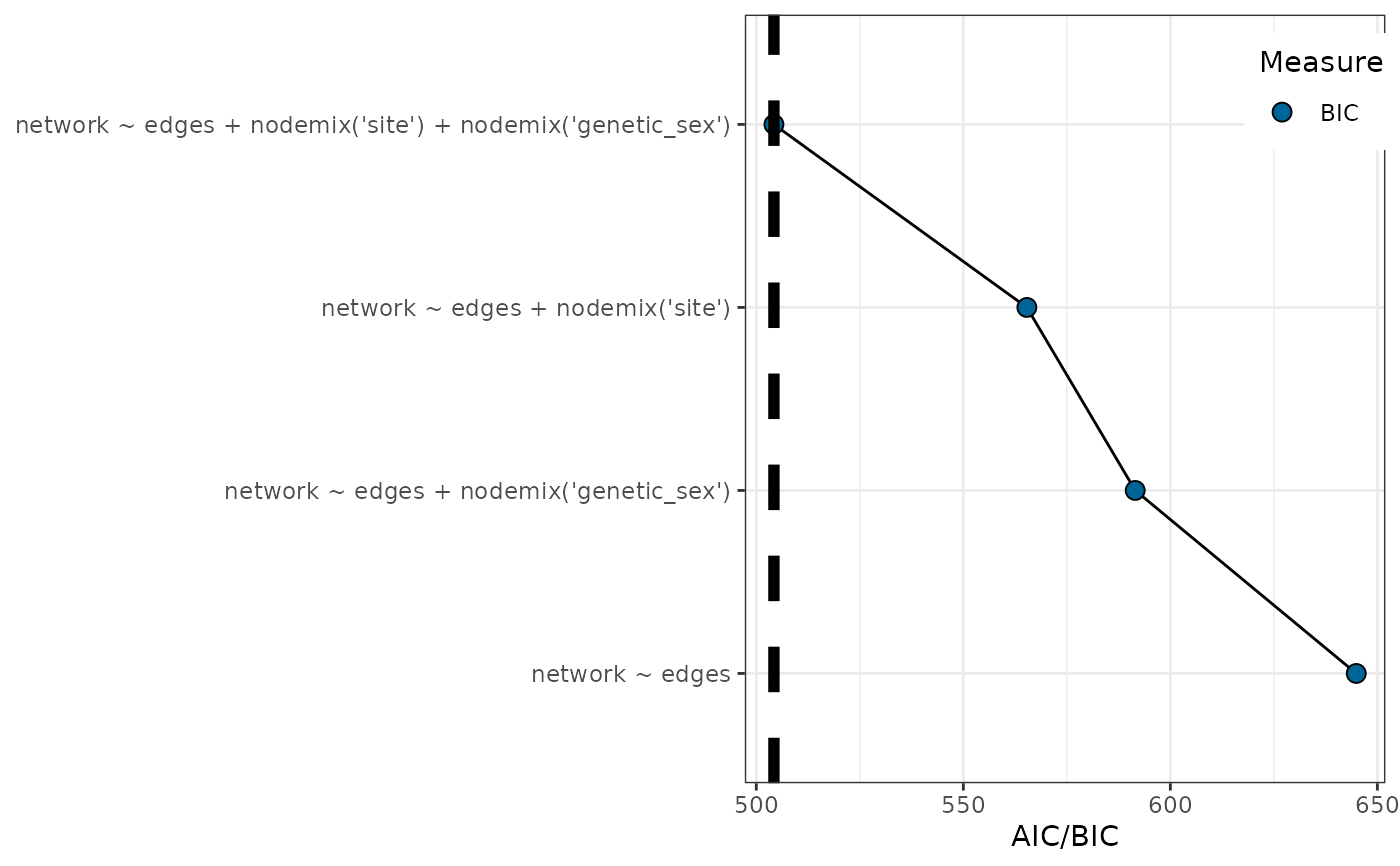
plot_ergm_bic(ergms) |> print()