A function to print the result of the combined analysis to screen
Source:R/printChASM.R
printChASM.RdA function to print the result of the combined analysis to screen
Examples
example_calls <- runChASM(rawReadCountsIn = example_data)
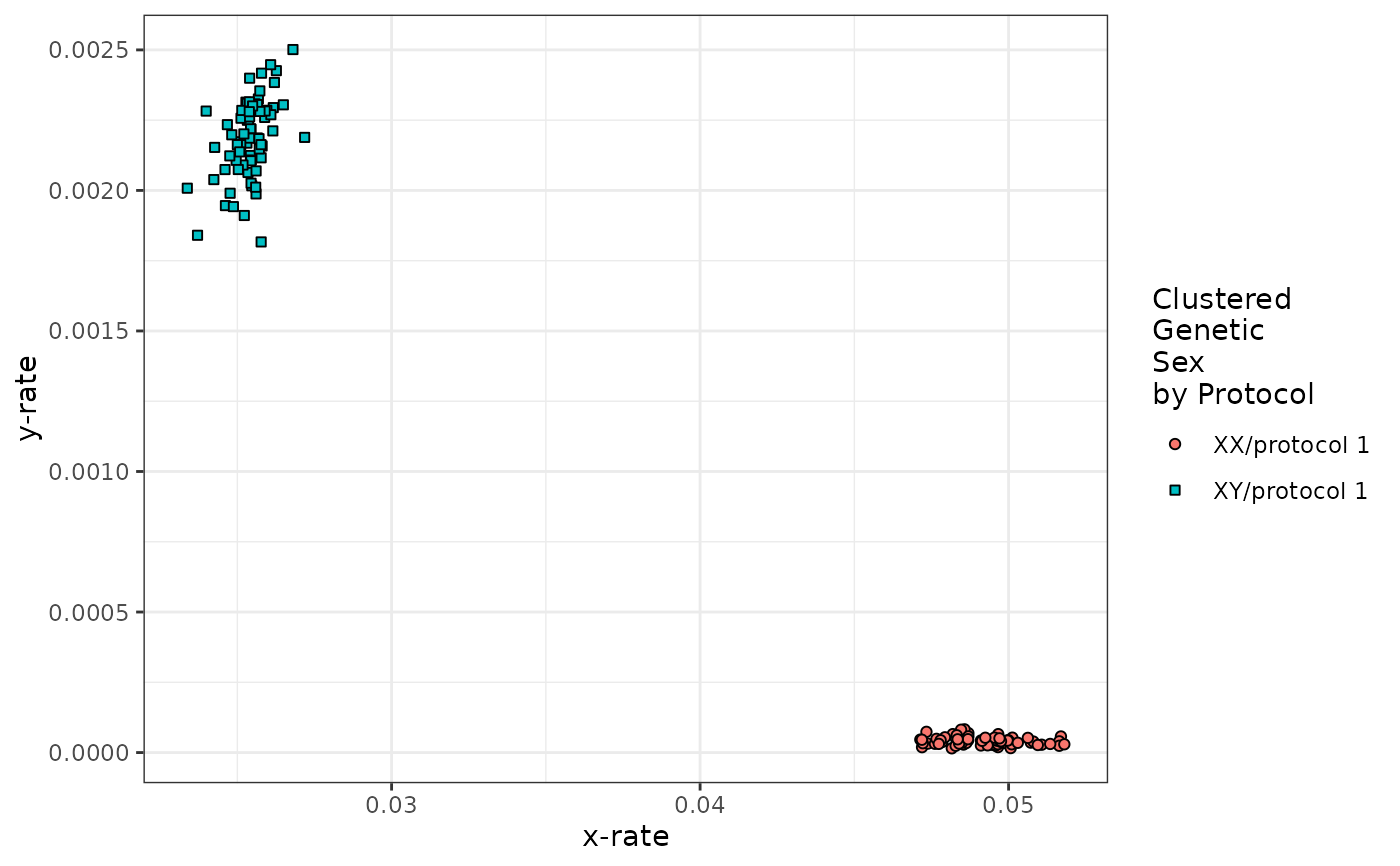 printChASM(inChASM = example_calls, lines = 10)
#> # A tibble: 222 × 11
#> sample protocol unusual flags autosomal_call sca_call C_call autosomal_total
#> <chr> <chr> <lgl> <int> <chr> <chr> <chr> <dbl>
#> 1 Ind_1_1 protoco… TRUE 3 No Aneuploidy XX No Si… 89539
#> 2 Ind_1_2 protoco… FALSE 1 No Aneuploidy XX No Si… 48376
#> 3 Ind_3_1 protoco… TRUE 2 No Aneuploidy XY No Si… 113335
#> 4 Ind_4_1 protoco… FALSE 0 No Aneuploidy XX No Si… 406472
#> 5 Ind_5_1 protoco… FALSE 0 No Aneuploidy XX No Si… 27731
#> 6 Ind_6_1 protoco… FALSE 0 No Aneuploidy XY No Si… 681256
#> 7 Ind_7_2 protoco… FALSE 0 No Aneuploidy XY No Si… 130928
#> 8 Ind_9_1 protoco… FALSE 0 No Aneuploidy XX No Si… 425906
#> 9 Ind_15… protoco… FALSE 1 No Aneuploidy XX No Si… 3364
#> 10 Ind_17… protoco… FALSE 1 No Aneuploidy XY No Si… 11699
#> # ℹ 212 more rows
#> # ℹ 3 more variables: sca_total <dbl>, automsomal_maxP <dbl>, sca_maxP <dbl>
printChASM(inChASM = example_calls, lines = 10)
#> # A tibble: 222 × 11
#> sample protocol unusual flags autosomal_call sca_call C_call autosomal_total
#> <chr> <chr> <lgl> <int> <chr> <chr> <chr> <dbl>
#> 1 Ind_1_1 protoco… TRUE 3 No Aneuploidy XX No Si… 89539
#> 2 Ind_1_2 protoco… FALSE 1 No Aneuploidy XX No Si… 48376
#> 3 Ind_3_1 protoco… TRUE 2 No Aneuploidy XY No Si… 113335
#> 4 Ind_4_1 protoco… FALSE 0 No Aneuploidy XX No Si… 406472
#> 5 Ind_5_1 protoco… FALSE 0 No Aneuploidy XX No Si… 27731
#> 6 Ind_6_1 protoco… FALSE 0 No Aneuploidy XY No Si… 681256
#> 7 Ind_7_2 protoco… FALSE 0 No Aneuploidy XY No Si… 130928
#> 8 Ind_9_1 protoco… FALSE 0 No Aneuploidy XX No Si… 425906
#> 9 Ind_15… protoco… FALSE 1 No Aneuploidy XX No Si… 3364
#> 10 Ind_17… protoco… FALSE 1 No Aneuploidy XY No Si… 11699
#> # ℹ 212 more rows
#> # ℹ 3 more variables: sca_total <dbl>, automsomal_maxP <dbl>, sca_maxP <dbl>