Data
data(skink, package = "monitoR")
skink |> glimpse()
#> Rows: 6,728
#> Columns: 4
#> $ time <dttm> 2024-03-25 14:34:01, 2024-03-25 14:35:01, 2024-03-25 14:36:…
#> $ X <dbl> 229, 248, 235, 250, 252, 248, 255, 256, 190, 141, 250, 261, …
#> $ Y <dbl> 213, 200, 221, 207, 214, 215, 275, 283, 370, 401, 243, 231, …
#> $ behaviour <chr> "Basking", "Basking", "Basking", "Movement", "Basking", "Bas…Processing
First we add a 4x4 grid
grid <- create_grid(range(skink$X), range(skink$Y), dim = c(4, 4))
skink_grid <- skink |> add_grid(grid)
plot_grid(grid, skink_grid)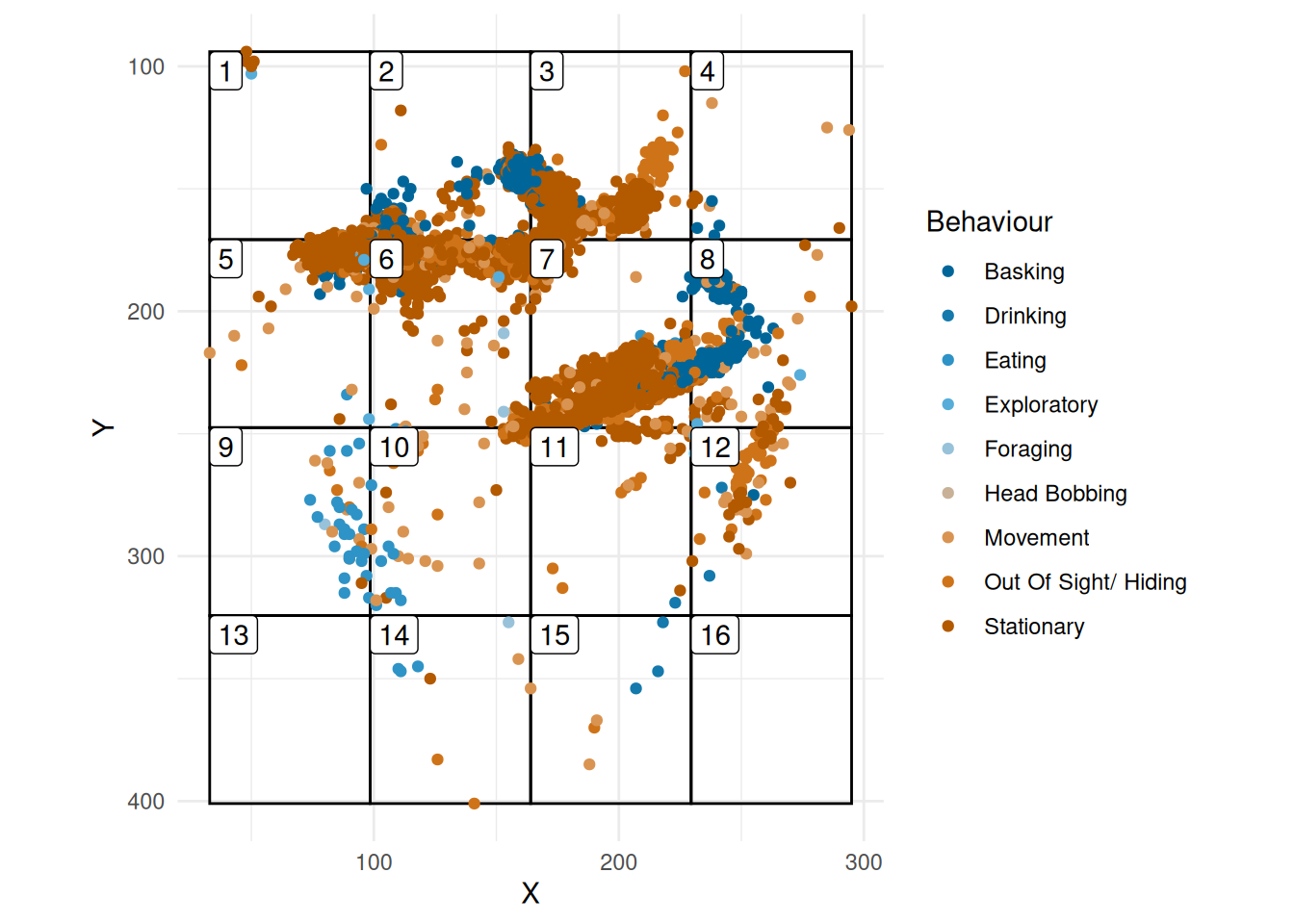
Index measures
Entropy
First we look at the entropy of observation for the data given in Figure 1.
calc_entropy(skink_grid$grid)
#> [1] 0.7944013Q1: Is it evenly spread? We know that the entopy for evenly spread data over a 4x4 grid using is
log10(16)
#> [1] 1.20412In our case, we have 0.7944013 which is smaller indicating non-even spread. We may wonder if this is significant?
To test this, we will create a empirical null distribution based on simulating data with equal spread of grids
null_entropy <- monitoR::get_entropy_null(
skink_grid$grid,
n_sims = 1000,
n_grid = 16
)If we look at the 95% percentiles
we see that it does not contain the observed value, so the observations are not evenly spread.
Entropy and behaviour
Next, we will calculate the entropy for each behaviour (Figure 2). In this we see a difference in entropy for different behaviours. For example, the most evenly spread is movement, while foraging is the least spread.
skink_grid |>
summarise(entropy = calc_entropy(grid), .by = behaviour) |>
mutate(
behaviour = fct_reorder(behaviour, entropy)
) |>
ggplot(aes(entropy, behaviour, fill = behaviour)) +
geom_col(show.legend = FALSE) +
harrypotter::scale_fill_hp_d("Ravenclaw") +
labs(x = "Entropy", y = NULL)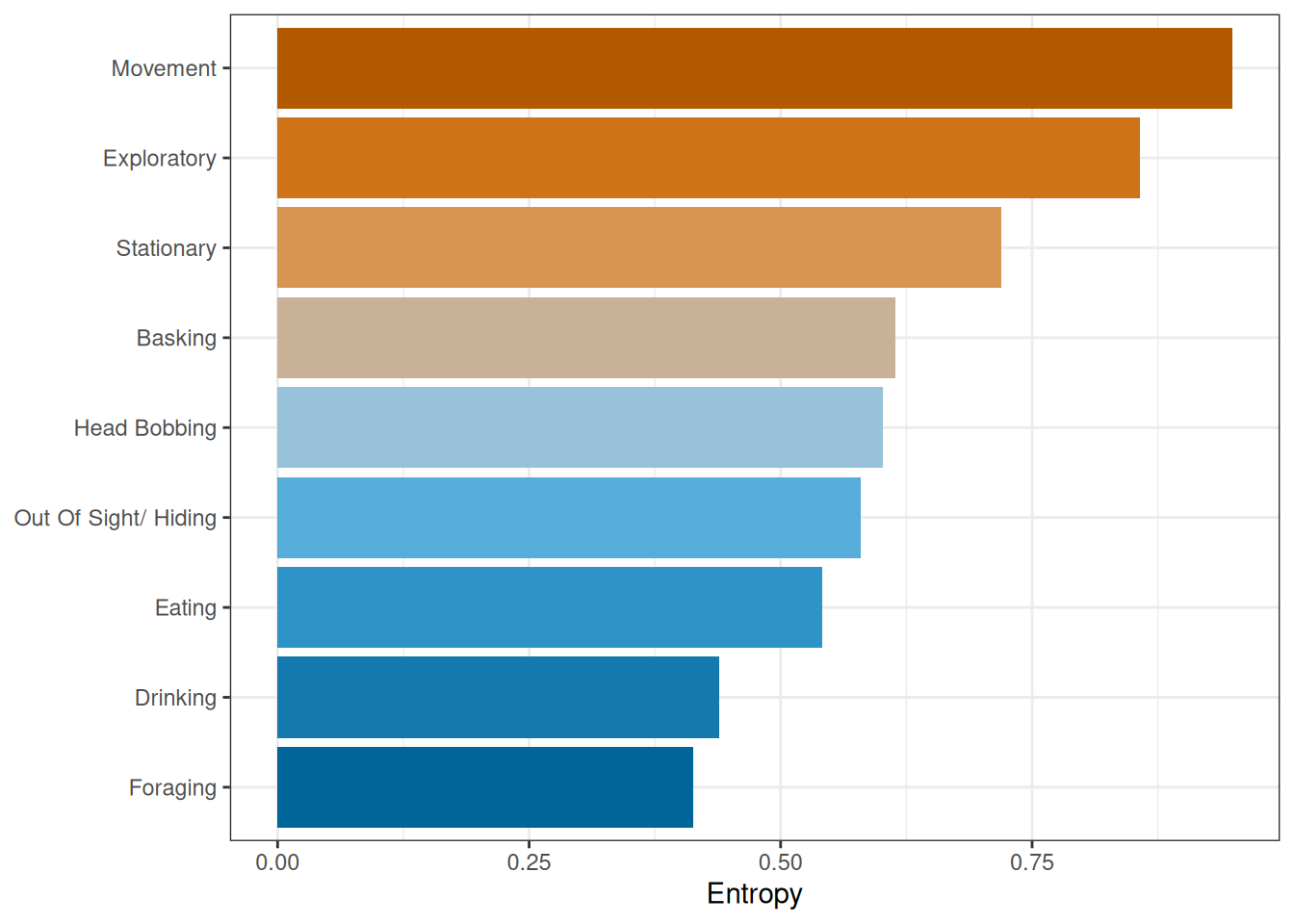
To assess the difference between the entropy for the different behaviours, we will used the squared distance of each entropy from the overall entropy
where is the entropy for the th behaviour, and is the entropy for all observations.
For Figure 2, we have
get_ess(skink_grid)
#> [1] 0.4849358As before, the question is
Is this significant?
To access this, we will use a permutation test, so we will
- Permute the grid recorded in the data
- Calculate ESS each time
ess_null <- get_ess_null(skink_grid, n_sims = 100)
quantile(ess_null, c(0.025, 0.975))
#> 2.5% 97.5%
#> 0.1483959 0.8317398So this time we find that the observed ESS is within the 95% empirical confidence interval and so is not surprising.
Environment effects
So, we will now focus on Basking (Figure 3). We will assume for illustration that there are rocks in Grid 5, 6, 7, 8, 2, and 3.
We want to ask the question of
Is it more likely to bask where there are rocks?
skink_bask <- skink_grid |>
filter(behaviour == "Basking")
plot_grid(grid, skink_bask)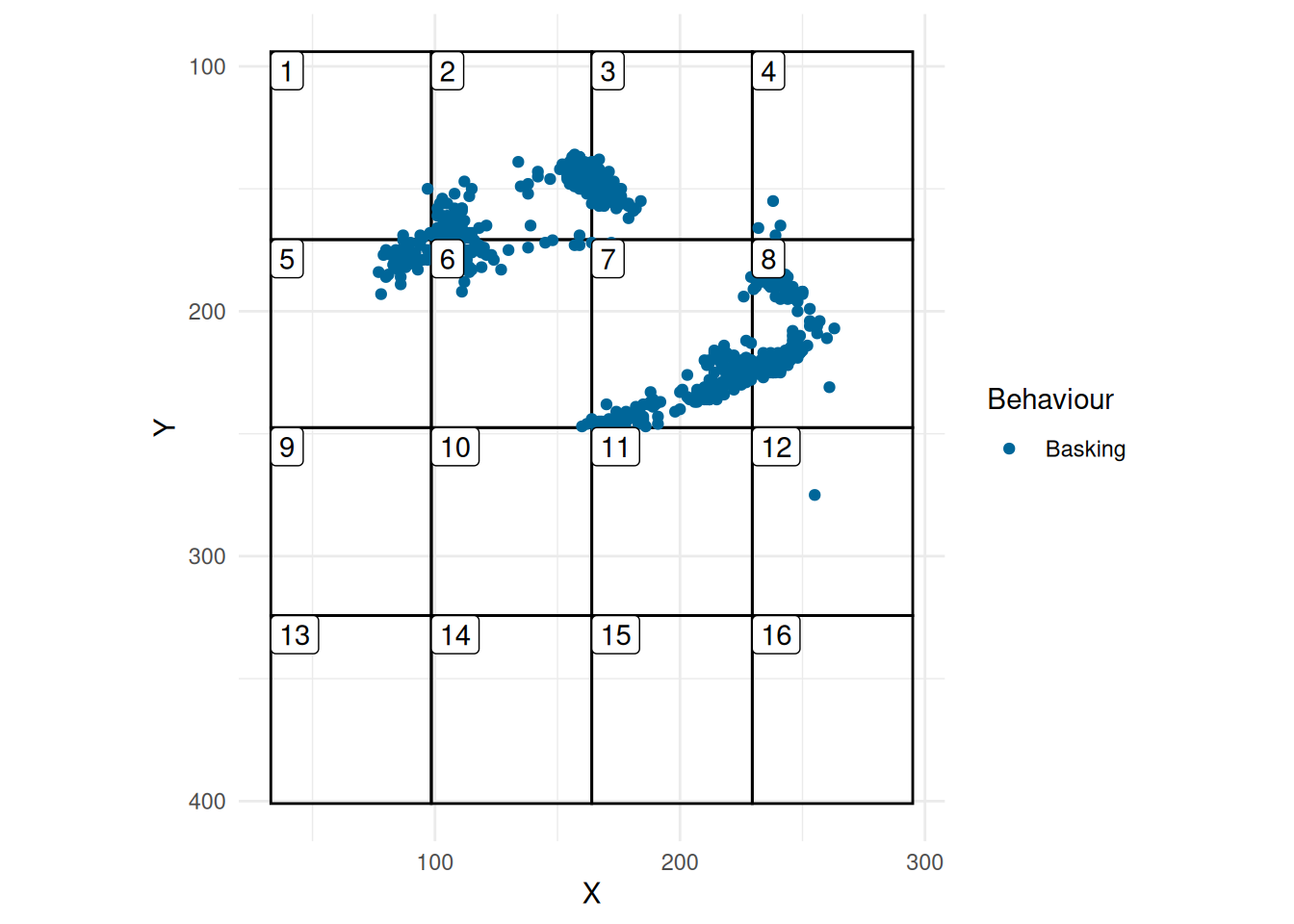
Table 1 gives the number of times that the skink is seen basking in each grid. As well, we have given the type of environment.
basking_df <- count_grid_behaviour(grid = grid, skink_bask)
basking_df <- basking_df |>
mutate(
environ = "normal"
)
basking_df$environ[c(2, 3, 5, 6, 7, 8)] <- "rocks"
basking_df |> gt()| grid | obs | environ |
|---|---|---|
| 1 | 5 | normal |
| 2 | 684 | rocks |
| 3 | 126 | rocks |
| 4 | 4 | normal |
| 5 | 36 | rocks |
| 6 | 47 | rocks |
| 7 | 199 | rocks |
| 8 | 166 | rocks |
| 9 | 0 | normal |
| 10 | 0 | normal |
| 11 | 7 | normal |
| 12 | 1 | normal |
| 13 | 0 | normal |
| 14 | 0 | normal |
| 15 | 0 | normal |
| 16 | 0 | normal |
We can fit a linear model of number of observations on environ, but we would also like to take into account the spatial dependency, that is, you will might be more likely to bask in grid next to one another. ?@fig-neigh show the neighbourhood of a corner grid, edge grid and a middle grid.
plot_neigh(grid, 1)
plot_neigh(grid, 2)
plot_neigh(grid, 10)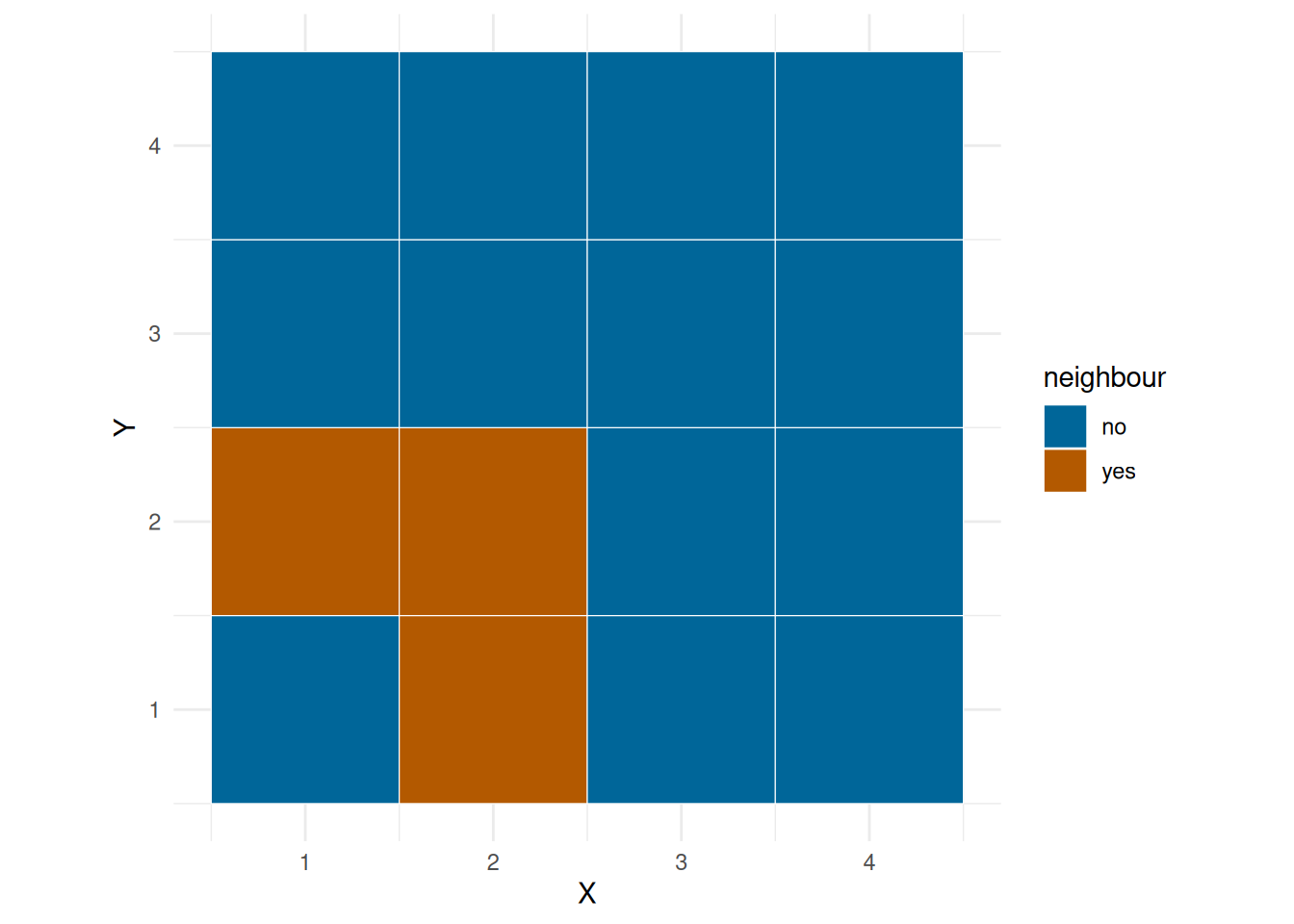
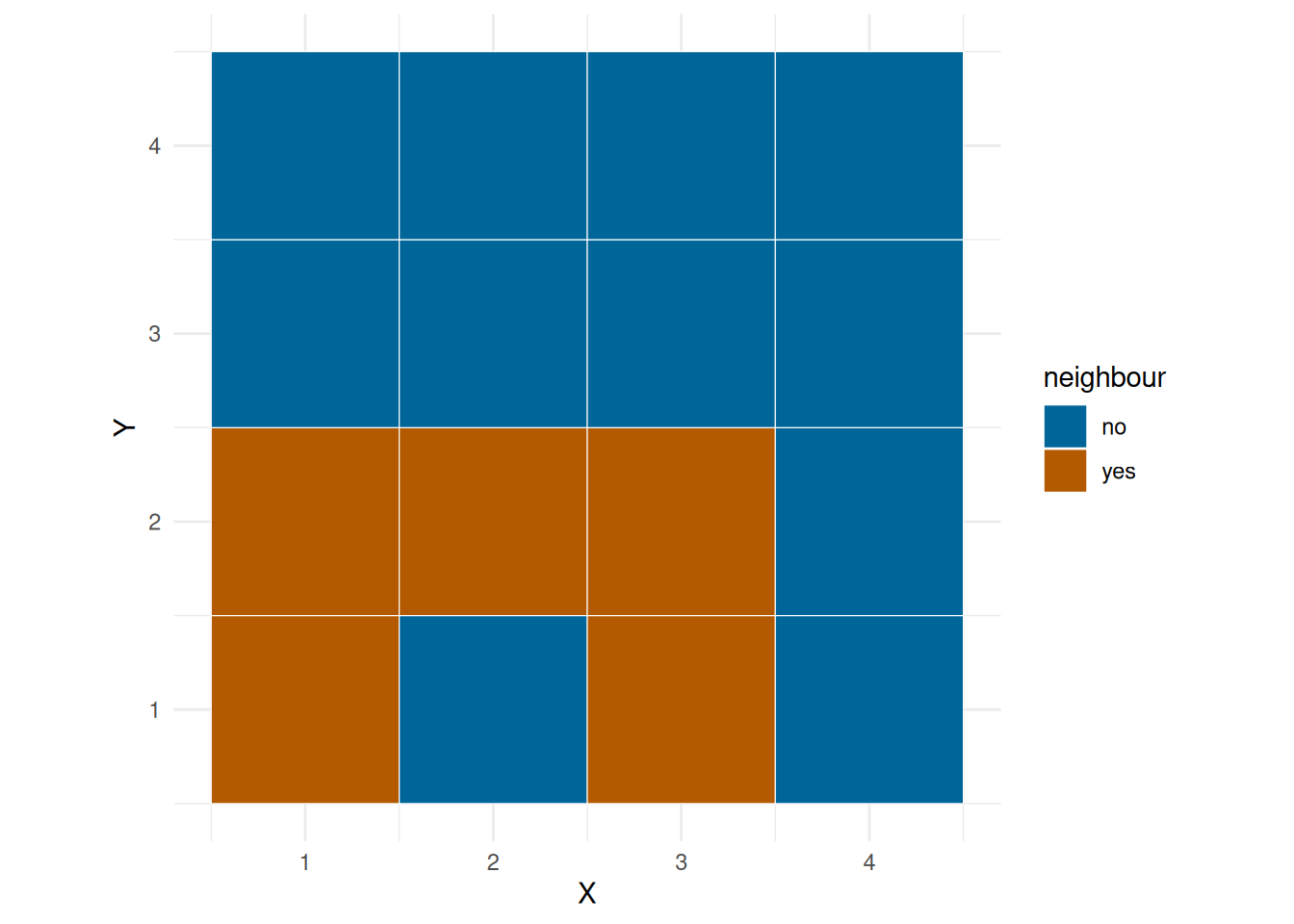
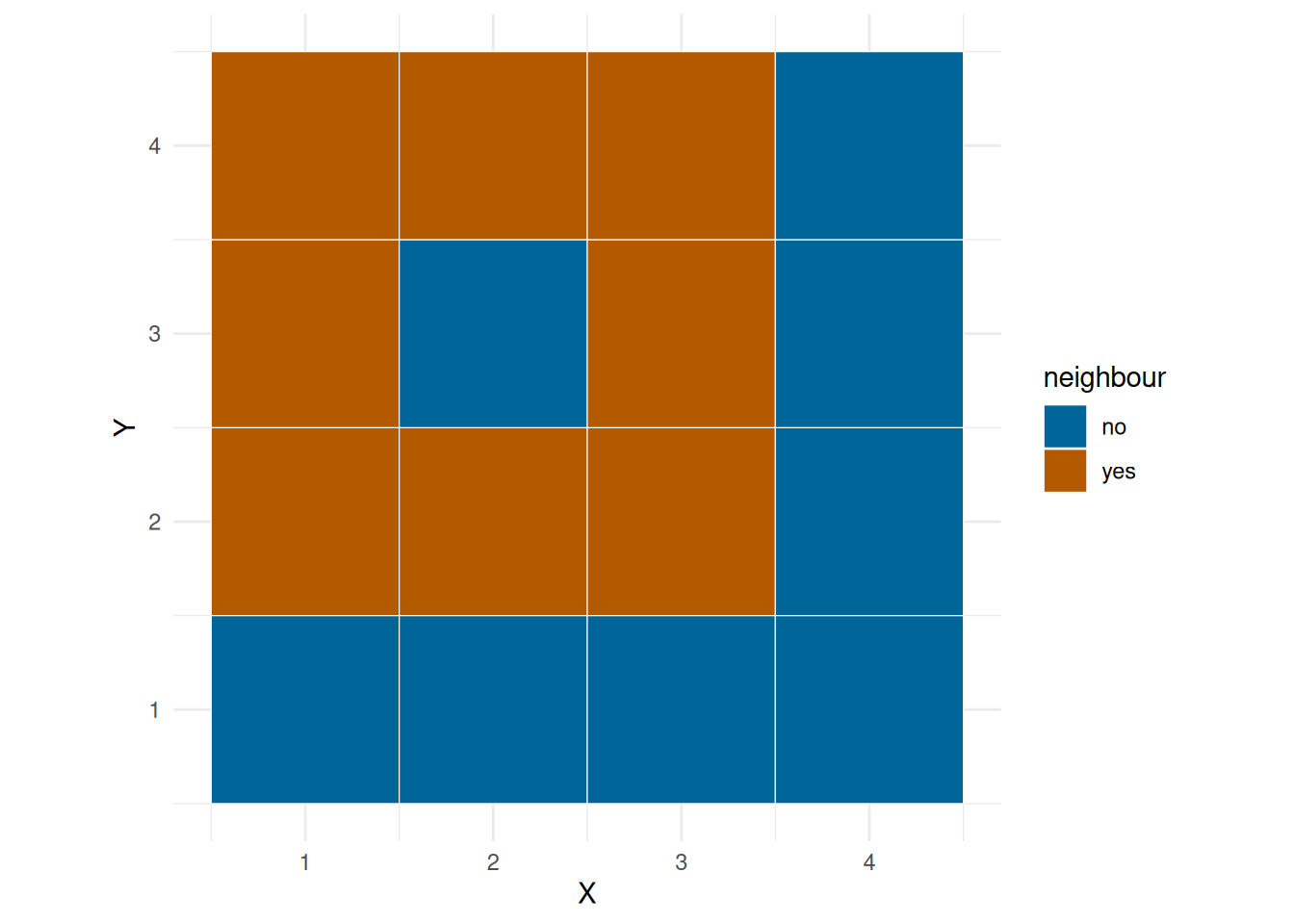
First we add the neigh mean.
M <- get_neigh_mat(grid)
basking_df$neigh <- as.numeric(M %*% basking_df$obs)
basking_df
#> # A tibble: 16 × 4
#> grid obs environ neigh
#> <dbl> <dbl> <chr> <dbl>
#> 1 1 5 normal 256.
#> 2 2 684 rocks 82.6
#> 3 3 126 rocks 220
#> 4 4 4 normal 164.
#> 5 5 36 rocks 147.
#> 6 6 47 rocks 132.
#> 7 7 199 rocks 129.
#> 8 8 166 rocks 67.4
#> 9 9 0 normal 16.6
#> 10 10 0 normal 36.1
#> 11 11 7 normal 51.6
#> 12 12 1 normal 74.4
#> 13 13 0 normal 0
#> 14 14 0 normal 1.4
#> 15 15 0 normal 1.6
#> 16 16 0 normal 2.67And then fit some models
M1 <- lm(obs ~ environ * neigh, data = basking_df)
summary(M1)
#>
#> Call:
#> lm(formula = obs ~ environ * neigh, data = basking_df)
#>
#> Residuals:
#> Min 1Q Median 3Q Max
#> -174.75 -3.78 -0.61 -0.30 375.19
#>
#> Coefficients:
#> Estimate Std. Error t value Pr(>|t|)
#> (Intercept) 0.4625 54.1866 0.009 0.9933
#> environrocks 481.9039 167.0120 2.885 0.0137 *
#> neigh 0.0205 0.5367 0.038 0.9702
#> environrocks:neigh -2.1217 1.2582 -1.686 0.1175
#> ---
#> Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
#>
#> Residual standard error: 137.3 on 12 degrees of freedom
#> Multiple R-squared: 0.5002, Adjusted R-squared: 0.3752
#> F-statistic: 4.003 on 3 and 12 DF, p-value: 0.03452
M2 <- update(M1, . ~ . - environ:neigh)
summary(M2)
#>
#> Call:
#> lm(formula = obs ~ environ + neigh, data = basking_df)
#>
#> Residuals:
#> Min 1Q Median 3Q Max
#> -167.30 -30.50 -20.25 2.68 457.09
#>
#> Coefficients:
#> Estimate Std. Error t value Pr(>|t|)
#> (Intercept) 23.7684 55.9865 0.425 0.6781
#> environrocks 233.3370 83.9025 2.781 0.0156 *
#> neigh -0.3655 0.5187 -0.705 0.4934
#> ---
#> Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
#>
#> Residual standard error: 146.8 on 13 degrees of freedom
#> Multiple R-squared: 0.3817, Adjusted R-squared: 0.2866
#> F-statistic: 4.013 on 2 and 13 DF, p-value: 0.04391
M3 <- update(M2, . ~ . - neigh)
summary(M3)
#>
#> Call:
#> lm(formula = obs ~ environ, data = basking_df)
#>
#> Residuals:
#> Min 1Q Median 3Q Max
#> -173.67 -18.92 -1.70 0.05 474.33
#>
#> Coefficients:
#> Estimate Std. Error t value Pr(>|t|)
#> (Intercept) 1.70 45.57 0.037 0.9708
#> environrocks 207.97 74.41 2.795 0.0143 *
#> ---
#> Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
#>
#> Residual standard error: 144.1 on 14 degrees of freedom
#> Multiple R-squared: 0.3581, Adjusted R-squared: 0.3123
#> F-statistic: 7.811 on 1 and 14 DF, p-value: 0.01433So it looks like there is no dependency.